WELCOME TO THE WEISS LAB
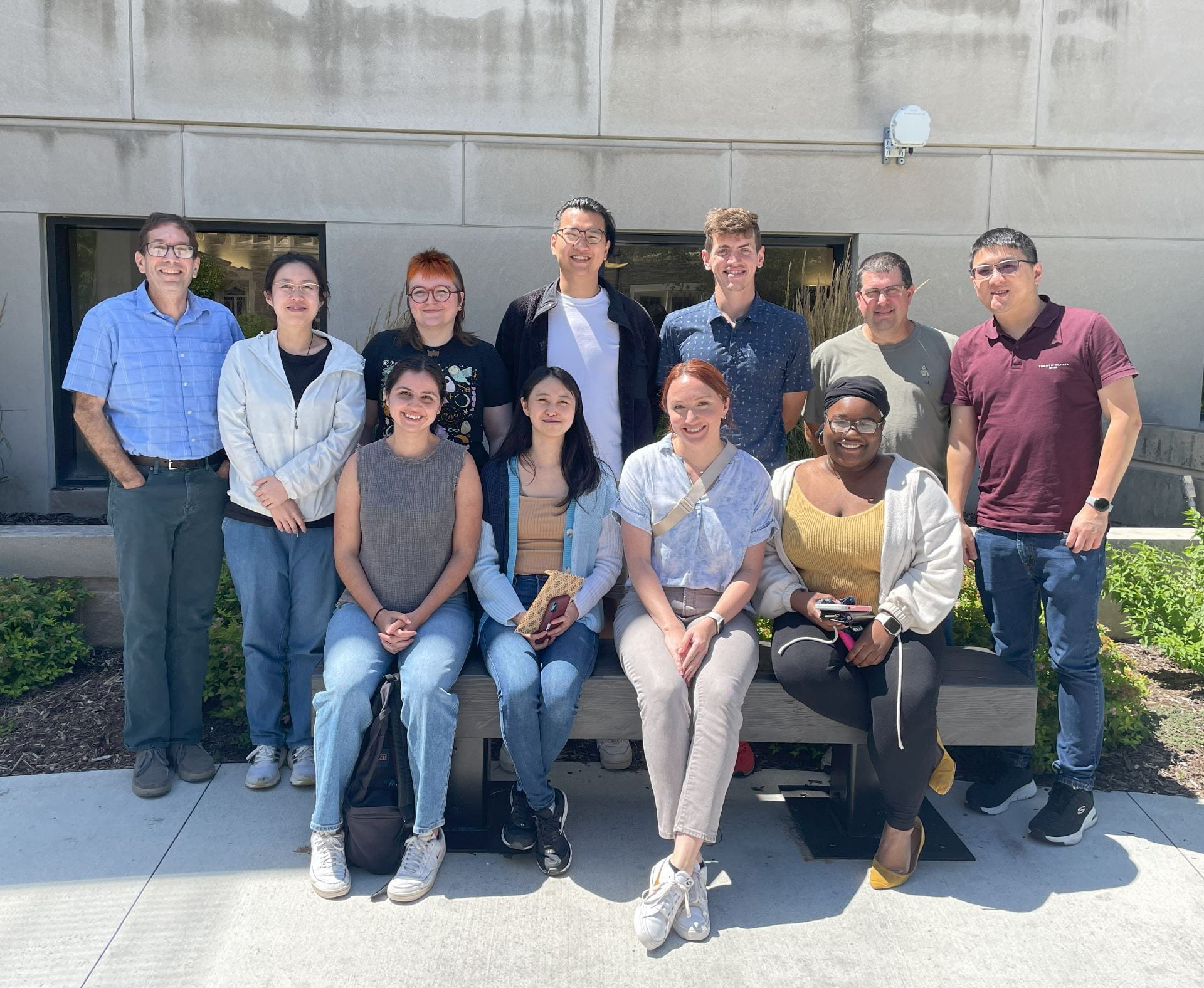


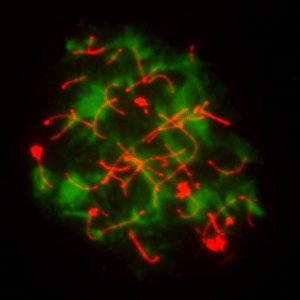
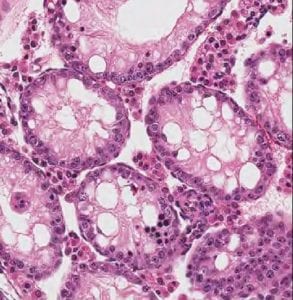
We are a lab in the Department of Biomedical Sciences at Cornell University. Our research focuses on how mammalian cells maintain genomic stability and how the loss of genomic integrity and metabolic control contributes to cancer and other diseases.
Check out our newest collaborative paper!
Check out our newest publication in eLife, a collaboration with the Cohen and Smolka labs: “A TOPBP1 allele causing male infertility uncouples XY silencing dynamics from sex body formation” (click here!)
Check out the Chronicle story featuring our research!: https://news.cornell.edu/stories/2024/03/sperm-formation-step-could-hold-clues-male-contraception
Congrats to Michelle on receiving a CVG Scholar Award!
The CVG Scholar Award is intended to recognize the genomics promise and exciting research goals of early stage trainees. With 26 applicants, the review was more competitive than ever before. Even with those odds, Michelle still received a top-3 score- congrats Michelle!

Check out our latest publications:
Bob takes on a new administrative role!
Bob takes on a new administrative role as Senior Associate Dean of the Graduate School (check out the article below!):
https://news.cornell.edu/stories/2023/08/robert-weiss-named-senior-associate-dean-graduate-school
Check out our newest preprint, as well as a collaborative study:
Amanda and co-authors publish a new preprint on miRNAs in germ cell tumors on bioRxiv:
The Weiss lab contributes to a new collaborative study on nanoparticle oral delivery:
Congratulations Andy on passing your A-Exam!



Kimaya and Victoria wrap up their summer research projects!
We sadly say goodbye to our two terrific summer students. Pictured below is Kimaya Bakhle, a VIP program participant and combined degree rotation student (jointly hosted with the Sethupathy lab). Kimaya was recently awarded the prestigious 2023 Boehringer Ingelheim Veterinary Research Award for Veterinary Students, and presented her research at the Veterinary Scholars Symposium.

Pictured below is Victoria Lopez-Scarim, a participant in the Molecular Biology & Genetics REU program. She recently gave a wrap up presentation about her summer research project, in which she investigated the 9-1-1 complex in collaboration with Gerardo.
Thanks to the both of you for all of your hard work!
Congratulations Michelle!
Congrats to Michelle on her successful A-exam!

Congratulations Dr. Mullmann!
Many congratulations to James on his successful defense! Good luck in vet school, we can’t wait to see what your future holds!


Weiss lab contributes to new collaborative publication
Check out our newest collaborative publication: A TOPBP1 Allele Causing Male Infertility Uncouples XY Silencing Dynamics From Sex Body Formation.
Bob and Destiny visit Spelman College
Destiny and Bob visit Spelman College to participate in the Spelman Career Fair and give a research seminar to the Spelman Research Initiative for Scientific Enhancement (RISE) program.


Congrats Gerardo on your NIH fellowship!

Congratulations, Dr. Fernandez!
Many congratulations to Irma on her successful defense, we can’t wait to see what you do next!



Many congratulations to Amanda on her successful PhD defense!!
![]()


Check out our newest collaborative publications!
Check out the Weiss Lab’s most recent collaborative publications!
Cancer Progression by Degrading the RNA Transcript Encoding a v-ATPase Subunit – https://pubmed.ncbi.nlm.nih.gov/36322753/
Welcome Justin and Jorlane!
A warm welcome to the Weiss lab’s summer ’22 students, Justin and Jorlane!!


Justin is a 3rd year biology major with a minor in environmental sciences at Morehouse College, visiting for the MBG REU program. Jorlane is a Puerto Rican rising senior majoring in Biomedical Sciences at the University of Puerto Rico in Ponce, and is here as a UPR-Ponce/CVM exchange student.
Congratulations Tanmay!!
Congratulations to Tanmay for graduating with a Bachelor’s in Human Biology, Health, and Society (HBHS)! We wish you much luck in all of your future endeavors.


Welcome Michelle! Plus a new publication:

A warm welcome to Michelle Liu! She has joined the Weiss lab as a graduate student in the field of Genetic Genomics & Development.
Also! Be sure to check out our newest paper, a collaboration with the Lin lab: Long-chain fatty acyl coenzyme A inhibits NME1/2 and regulates cancer metastasis: https://pubmed.ncbi.nlm.nih.gov/35259022/ .
Weiss Lab contributes to a new paper published in Stem Cell Reports from the Shioda group at MGH
Check out the paper here:
Expanding homogeneous culture of human primordial germ cell-like cells maintaining germline features without serum or feeder layers. https://www.cell.com/stem-cell-reports/fulltext/S2213-6711(22)00056-X
Weiss & Smolka labs publish back to back papers in eLife!
Congratulations to Catalina Pereira, Gerardo A. Arroyo-Martinez, and the other authors involved! Check out the papers here:
Multiple 9-1-1 complexes promote homolog synapsis, DSB repair, and ATR signaling during mammalian meiosis. https://doi.org/10.7554/eLife.68677.sa0
Phosphoproteomics of ATR signaling in mouse testes. https://doi.org/10.7554/eLife.68648
Weiss Lab Holiday Social!


Members of the Weiss lab celebrated the 2021 holiday season with brunch from Agava!
Congratulations Destiny and Gerardo!!
Congratulations to both Destiny Van and Gerardo A. Arroyo Martinez on passing their A-exams this summer!!


Check out these BioRxiv preprints involving the Weiss Lab!
IGF2BP2 Promotes Cancer Progression by Degrading the RNA Transcript Encoding a v-ATPase Subunit: https://doi.org/10.1101/2021.04.01.438101
Multiple 911 complexes promote homolog synapsis, DSB repair and ATR signaling during mammalian meiosis: https://doi.org/10.1101/2021.04.09.439198
Phosphoproteomics of ATR Signaling in Prophase I of Mouse Meiosis: https://doi.org/10.1101/2021.04.13.439649
Building a Bridge between Benchside and Bedside: A reflection by Ravi Dhawan and coauthors
Cancer Researchers and Cancer Patients Communicating about Basic Science
Our collaborations have inspired us to translate the complexities of cancer research into accessible scientific narratives.
Read the full article here: https://cancercommunity.nature.com/documents/cancer-researchers-and-cancer-patients-communicating-about-basic-science-521d1eb4-1066-4e9e-92a6-1bea37062e1a
Congratulations to Irma Fernandez, James Mullmann, and Ravi Dhawan on their publication in Oncogene!
Collaborative paper with the Cerione and Lin labs: https://www.nature.com/articles/s41388-020-01637-w
I’m dreaming of a [Weiss] Christmas (and all other holidays!)
Trust me, I’m the Doctor: Welcome Dr. Pereira!
Cat’s dissertation is formally approved. Congratulations to Dr. Pereira!
Congratulations, Irma Fernandez!
Irma Fernandez is selected as an Edward A. Bouchet Graduate Honor Society Scholar! This highly selective honor recognizes graduate students for outstanding scholarly achievement and efforts to promote diversity.
Undergraduate Grants
Ravi Dhawan is awarded a grant from the Dextra Undergraduate Research Endowment Fund, and Anabella Maria Galang is awarded a grant from the Ann S. and Robert R. Morley Student Research Fund. Congratulations!
Welcome BMCB PhD Student: Destiny Van!
“It is not in the stars to hold our destiny but in ourselves.” ~ William Shakespeare
The Weiss Lab welcomes Destiny Van!
James Mullmann passes his A-Exam
Congratulations to James Mullmann on passing his A-exam!
Parting is such sweet sorrow: Matthew Guo and Michael Downey graduate
Michael Downey and Matthew Guo have receive their undergraduate degrees from Cornell and are moving on to the next stages in their lives.
James Mullmann awarded NIH F30 Fellowship
James Mullmann has been awarded an NIH F30 Fellowship for his project “Elucidating the tumor suppressive effects of the sirtuin, SIRT1, in triple-negative breast cancer.” Congratulations, James!
Amanda Loehr passes her A-exam
Congratulations to DVM/PhD student Amanda Loehr on passing her A-exam!
Congratulations to Amanda for being awarded an NIH F30 fellowship!
Amanda received an NIH F30 fellowship for her project titled “The role of the DNA damage response in the development and therapeutic sensitivity of malignant testicular germ cell tumors.”
Congratulations to Weiss lab members for their new publications!
Weiss lab holiday lunch at the Statler!
Congratulations to Ravi Dhawan for his new research grant from the Dextra Undergraduate Research Endowment Fund!
Congratulations to Matthew Guo for presenting his thesis project at the Fall 2019 CURB symposium and being awarded as First Prize: Best in Biology!
Weiss lab members, summer 2019
What a summer it’s been! This summer, we welcomed Suzin Webb as our new lab manager! We are so happy to have you. We also had three wonderful summer students: Carla Patricia Reyes-Flores, Natrine Cheuk, and Jamie Cockey. We are so glad to have had you in the lab this summer, and we look forward to hearing what great things you accomplish in the future!
Carla and Matthew present their research at the Summer Institute for Life Sciences Research Symposium!
Congratulations to MBG-REU student Carla Patricia Reyes Flores for her outstanding end-of-summer poster presentation!
Congratulations to Irma Fernandez for being selected for a 2019 Gilliam Fellowship for Advanced Study from HHMI!
See the Cornell news article here: https://www.vet.cornell.edu/news/20190903/graduate-student-irma-fernandez-wins-award-howard-hughes-medical-institute. See the HHMI announcement here: https://www.hhmi.org/news/44-gilliam-fellowships-awarded-to-support-diversity-and-inclusion-in-science
Congratulations to Amanda Loehr and co-authors on their new review in Andrology
Bloom, J.C., Loehr, A.R., Schimenti, J.C., and Weiss, R.S. (2019) Germline genome protection: implications for gamete quality and germ cell tumorigenesis.Andrology 7:516-526. Pubmed.
Bob and Bob publish a perspective piece on the Cornell Cancer Partnership in Cancer Research
Riter, R.N. and Weiss, R.S. (2019) Connecting students with patients and survivors to enhance cancer research training. Cancer Research, in press.
The Weiss lab partners with the Cerione and Lin groups to receive a new R01 from the NIH!
https://research.cornell.edu/research/cancer-and-sirt5
Congratulations to Brenna Remick for graduating with honors in biological sciences! We will miss you! Cannot wait to hear the great things you will accomplish at Berkeley!
Our lab attends the 23rd Annual Buffalo DNA Replication & Repair Symposium
Thank you to the University at Buffalo Jacobs School of Medicine and Biomedical Sciences and the conference organizers for a wonderful conference! Catalina, PhD candidate, presented a talk titled “9-1-1 DNA Damage Response Clamp Subunit RAD9A Promotes Checkpoint Signaling to Maintain Genomic Stability,” and Matthew and Michael, undergraduates, presented a poster titled “Multiple Mammalian 9-1-1 DNA Damage Response Complexes Are Important for Meiotic Chromosome Maintenance.”
Congratulations to Matthew Guo for his successful oral presentation at the National Conference of Undergraduate Research!
Weiss lab lunch over spring break to say farewell to Dr. Elizabeth Moore! Best of luck at your post-doc in the Fischbach lab!
PhD student Irma Fernandez is featured by the Cornell Office of the Vice Provost for Research
Sirtuins’ connection to cancer is a big topic in life sciences research. What are they and why are they important to the hunt for cancer therapies? Read more about the work Irma in our lab here: https://research.cornell.edu/news-features/whats-all-talk-about-sirtuins
Congratulations to Dr. Elizabeth Moore for her successful dissertation defense!
The Weiss and Smolka labs have been awarded a collaborative NIH research grant!
Infertility and birth defects often arise due to improper genetic quality control during meiosis. We are working with Marcus Smolka’s lab in Molecular Biology and Genetics to resolve how the cellular DNA damage response ensures genetic quality control during meiosis and enables the efficient and accurate production of gametes. Read the article here: https://research.cornell.edu/research/genetic-quality-control-during-meiosis
Congratulations to Dr. Elizabeth Moore for her second place finish for the prestigious 2018 Young Investigator Award (YIA) sponsored by the American Veterinary Medical Association (AVMA) and the American Veterinary Medical Foundation (AVMF).
Congratulations to Naomi for her successful poster presentation!
Congratulations to Irma Fernandez for passing her A-exam!
Congratulations to Darshil Patel on his successul PhD defense
Congratulations to Matthew Guo and Michael Downey on their first place prize at the Cornell Undergraduate Research Board Spring Forum for their poster “Roles of the Mammalian 9-1-1 DNA Damage Response Clamps in Meiotic Chromosome Maintenance.”
Darshil and Irma present at annual cancer symposium
Congratulations to Tim, Amy, and co-authors on their new paper in Cell Reports
http://www.cell.com/cell-reports/fulltext/S2211-1247(17)31541-3
Study Reveals Why Testicular Cancer Responds to Chemo
http://news.cornell.edu/stories/2017/11/study-reveals-why-testicular-cancer-responds-chemo
Dr. Robert Weiss Appointed as New Associate Dean for Research and Graduate Education
https://www2.vet.cornell.edu/news/20171009/dr-robert-weiss-appointed-new-associate-dean-research-and-graduate-education
The Weiss and Richards Labs get together for a wine and design outing.
Weiss lab retreat at Fillmore Glen State Park
Farewell to new Cornell graduates, Ellen Hong (BS 2017) and Qiming Jin (MEng 2017)
Weiss lab tours the Ithaca Beer Company
Weiss lab contributes to collaborative research with the Reinhart-King lab on the effects of matrix stiffness on tumorigenesis.
http://www.news.cornell.edu/stories/2017/01/softening-tumor-tissue-could-aid-cancer-drug-delivery
Congratulations to Tim Pierpont on his successful PhD defense
Congratulations to Catalina Pereira for passing her A-exam!
Congratulations to Darshil Patel for his outstanding poster presentation award at the 2016 Genomic Instability GRC in Hong Kong
“Tumor promoting functions for the DNA damage response protein HUS1”
Congratulations to Elizabeth Moore for her best poster award at the 2016 Cornell BBS symposium
“A novel role for the Fanconi Anemia pathway in bile acid homeostasis”
The Weiss lab participates in the Ithaca Cancer Moonshot Summit
http://www.news.cornell.edu/stories/2016/07/local-center-part-biden-s-cancer-moonshot-summit
Congratulations to Charlton Tsai and Zanah Francis, Cornell graduates in the class of 2016!
Charlton Tsai is named a 2016 Merrill Presidential Scholar
http://www.news.cornell.edu/stories/2016/05/33-seniors-honored-2016-merrill-presidential-scholars
The Weiss lab collaborates with the Hening Lin lab (Dept. of Chemistry and Chemical Biology) to publish new papers on Sirt2 and Sirt5
Congratulations to Darshil Patel for passing his A-Exam!!!
Congratulations to Joanna Mleczko on her successful B-exam!
Congratulations to Tim Pierpont for taking second place at the 2015 Cornell Stem Cell Symposium poster competition!
INVESTIGTATING HOW THE DNA DAMAGE RESPONSE AND STEM CELL PROPERTIES OF TESTICULAR GERM CELL CANCERS INFLUENCE CHEMOSENSITIVITY
Pierpont, Timothy M.1, Lyndaker, Amy M.1, Anderson, Claire M.1, Roden, Jamie L.1, Bagepalli , Lina1, Kataria, Nandita1, Braxon, Alicia1 , Hu, Hilary1, Garness, Jason2, Cook, Matthew S.2, Capel, Blanche2, Schlafer, Donald H.1, Southard, Teresa1, and Weiss, Robert S1.
1Department of Biomedical Sciences, College of Veterinary Medicine, Cornell University, Ithaca, NY, USA and 2Department of Cell Biology, Duke University Medical Center, Durham, NC, USA
Congratulations to Dr. Robert Weiss on being promoted to full professor!
Congratulations to Gabi Balmus and co-authors!
Congratulations to Gabi Balmus and co-authors on the acceptance of their manuscript: “HUS1 regulates in vivo responses to genotoxic chemotherapies” for publication in Oncogene.
Congratulations to Pei Xin Lim and co-authors!
Congratulations to Pei Xin Lim and co-authors on acceptance of their manuscript: “Genome Protection by the 9-1-1 Complex Subunit HUS1 Requires Clamp Formation, DNA Contacts, and ATR Signaling-Independent Effector Functions” for publication in the Journal of Biological Chemistry.